Lorenzo-Orts Lab
Research Publications Group Members BiographyMaternal mRNA regulation
Gene expression ultimately depends on translation. Although mRNAs serve as templates for protein production, mRNA and protein levels often correlate poorly. Translational regulation contributes to this uncoupling and provides a powerful mechanism for rapid, precise and dynamic control of protein output across multiple biological contexts.
One such context is early embryonic development, during which gene expression is governed at the post-transcriptional level through the regulation of maternal mRNAs. These mRNAs are synthesised during oogenesis and stored in the mature oocyte, where their translation is tightly regulated in both time and space. This strategy enables swift modulation of protein synthesis without the need for de novo transcription, ensuring robust maternal control over developmental programmes during the earliest stages of life.
Translational repression during early development
As maternal mRNAs are stockpiled in the mature oocyte, robust translational repression mechanisms must be established to (1) conserve cellular energy and (2) prevent premature protein synthesis, which could be detrimental to embryonic development. During my postdoctoral work, I identified two distinct mechanisms that repress translation at the level of ribosomes and mRNAs (Figure 1).

Ribosomes are maternally deposited to support protein synthesis during early embryogenesis. How ribosomes are stored in an inactive state within mature oocytes has long remained unclear. We discovered that maternal ribosomes in fish and frogs are bound by four factors that block functionally important sites, thereby regulating both ribosome abundance and activity during early development.
In the case of maternal mRNAs, translational repression is linked to deadenylation. While deadenylation triggers decapping and subsequent mRNA degradation in somatic cells, maternal mRNAs with short polyadenine tails remain stable in the embryo. We found that the germline-specific protein eIF4E1b binds the 5′ mRNA cap and promotes the storage and repression of maternal mRNAs in processing bodies (P-bodies). Although eIF4E1b is a paralogue of the translation initiation factor eIF4E, its repurposed role as a translational repressor rather than an activator highlights the unique regulatory landscape of gene expression in the vertebrate germline.
Translational activation
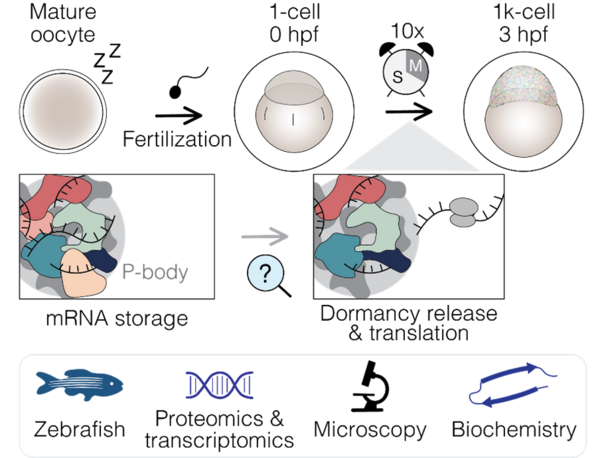
After fertilisation, translation gradually increases to meet the demand for new proteins in the embryo. As maternal mRNAs are bound by proteins involved in translational repression and stored in P-bodies, their translation likely requires remodelling and potential relocalisation of these mRNAs to the cytosol. While several mechanisms underlying the translational repression of maternal mRNAs have been described (see above), much less is known about their translational activation.
How is the embryonic proteome remodelled to transition from translational repression to initiation? What molecular complexes drive the activation of maternal mRNAs? How is translation initiated, and when do these events take place? These are some of the questions our newly established lab is currently addressing (Figure 2).
Development and beyond
Controlling the emergence of proteins at the correct time and place is essential for development. While early embryos offer a major advantage for studying post-transcriptional gene regulation due to the absence of transcription, the mechanisms uncovered here are likely to operate in other contexts, including ageing and disease. During ageing, there is a gradual uncoupling of mRNA and protein levels, partly driven by changes in translational activity. Moreover, dysregulation of mRNA metabolism is associated with diseases such as leukaemia and Parkinson’s disease. We use embryos to uncover the mechanisms underlying mRNA regulatory processes that are relevant for development and beyond.
Approaches
Our lab combines in vivo and in vitro approaches to study maternal mRNA regulation across multiple levels. Zebrafish is our primary in vivo model, offering powerful genetic tools and abundant embryos for applications such as proteomics, microscopy and transcriptomics. We complement these studies with in silico modelling approaches (e.g. AlphaFold), protein biochemistry and structural biology to link molecular mechanisms to their roles in development.
Recruitment
We are driven by scientific curiosity and hypothesis-based research. We welcome students and postdocs who are excited about uncovering new principles of early embryonic development and mRNA regulation using diverse experimental approaches. If you are interested, please get in touch!